CRH-221・229
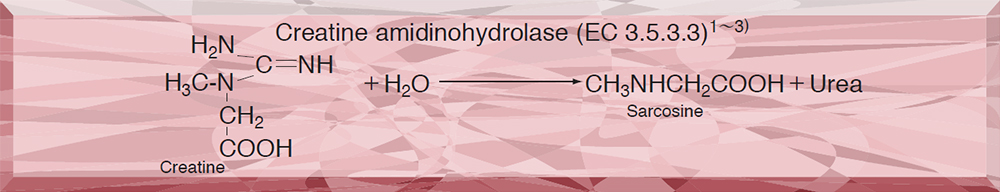
CREATINE AMIDINOHYDROLASE from Microorganism
PREPARATION and SPECIFICATION
| Appearance | (-221) White amorphous powder, lyophilized (-229) Clear solution |
|
|---|---|---|
| Activity | GradeⅡ (-221) 4.0U/mg-solid or more (-229) 2,000U/ml or more |
|
| Contaminants | NADH oxidase | ≤5.0×10-2% |
| Catalase | ≤2.0% | |
| Stabilizers | Sugars, EDTA | |
PROPERTIES
| Stability | (-221)Stable at −20℃ for at least one year (-229) Stable at 4℃(Fig.1) |
|---|---|
| Molecular weight | approx. 67,000 (by gel filtration) |
| Structure | 2 subunits per enzyme molecule |
| Isoelectric point | 4.5±0.1 |
| Michaelis constant | 4.5×10-3M (Creatine) |
| Inhibitors | Hg++,Cu++Ag+,SH reagent (NEM), PCMB |
| Optimum pH | 6.5-7.5(Fig.2) |
| Optimum temperature | 40-50℃(Fig.3) |
| pH Stability | pH 4.0−10.0 (25℃, 20hr)(Fig.4) |
| Thermal stability | below 50℃ (pH 7.5, 30min)(Fig.5) |
| Effect of various metals | (Table 1) |
APPLICATIONS
This enzyme is useful for enzymatic determination of creatine and creatinine when coupled with creatinine amidohydrolase (CNH-211, CNH-311) and sarcosine oxidase (SAO-351) in clinical analysis.
ASSAY
Principle

The appearance of yellow dye formed by condensation of urea and p-dimethylaminobenzaldehyde (DAB) (Ehrlich reaction) is measured at 435nm by spectrophotometry.
Unit definition
One unit causes the formation of one micromole of yellow dye per minute under the conditions described below.
Method
Reagents
| A. Creatine solution | 0.1M [1.49g creatine (Nacalai tesque)/100ml of 50mM phosphate buffer, pH 7.5](Prepare freshly) |
|---|---|
| B. DAB solution | Dissolve 2.0g of DAB in 100ml of dimethylsulfoxide and, to this solution, add 15ml of conc. HCl solution. |
| C. Enzyme diluent | 50mM Phosphate buffer, pH 7.5 |
Procedure
1.Pipette 1.0ml of the substrate solution (A) into a test tube and equilibrate at 37℃ for about 5 minutes.
| Concentration in assay mixture | |
|---|---|
| Phosphate buffer | 50 mM |
| Creatine | 90 mM |
2.Add 0.1ml of the enzyme solution* and mix.
3.After exactly 10 minutes at 37℃, add 2.0ml of DAB solution (B) to stop the reaction.
4.Incubate at 25℃ for 20 minutes.
5.Measure the optical density at 435nm against water (OD test).
At the same time, prepare the blank by first mixing the substrate solution with 2.0ml of DAB solution after 10 min-incubation at 37℃, followed by addition of the enzyme solution, and carry out the same procesure as test (procedure 4 and 5)(OD blank).
*Dissolve the enzyme preparation in ice-cold enzyme diluent (C) and dilute to 2.0-3.0U/ml with the same buffer, immediately before assay.
Calculation
Activity can be calculated by using the following formula :
Volume activity (U/ml) =
-
ΔOD (OD test−OD blank)×Vt×df
0.321×1.0×t×Vs
= ΔOD×9.65×df
Weight activity (U/mg) = (U/ml)×1/C
| Vt | : Total volume (3.1ml) |
| Vs | : Sample volume (0.1ml) |
| 0.321 | : Millimolar extinction coefficient of yellow dye(cm2/micromole) |
| 1.0 | : Light path length (cm) |
| t | : Reaction time (10 minutes) |
| df | : Dilution factor |
| C | : Enzyme concentration in dissolution (c mg/ml) |
REFERENCES
1)D.Tsuru; Nucleic Acid and Amino Acids, 35, 31 (1977).
2)T.Yoshimoto, I.Oka and D.Tsuru; Arch.Biochem.Biophys., 177, 508 (1976).
3)T.Yoshimoto ,I.Oka and D.Tsuru; J.Biochem., 79, 1381 (1976).
4)D.Tsuru; Rinsho Kensa, 22, 1331 (1978).
Table 1. Effect of Various Chemicals on Creatine amidinohydrolase
[The enzyme dissolved in 50mM Tris-HCl buffer, pH 7.5 (80U/ml) was incubated with each chemical at 25℃ for 30 minutes.]
-
Chemical Concn.(mM) Residual activity(%) None ー 100 Metal salt 1.0 CaCl2 102 MnCl2 96 MgCl2 103 NiCl2 104 CoCl2 104 Ba(OAc)2 98 Cd(OAc)2 98 FeCl3 89 FeSO4 81 HgCl2 0 ZnSO4 63 CuSO4 0 Pb(OAc)2 98 AgNO3 0 EDTA 20 98 α,α′-Dipyridyl 1.0 102 -
Chemical Concn.(mM) Residual activity(%) NaF 1.0 99 PCMB 0.33 0 MIA 1.0 94 IAA 1.0 100 NaN3 10 96 o-Phenanthroline 1.0 99 Hydroxylamine 1.0 99 NEM 10 32 Triton X-100 0.5% 99 Brij 35 0.5% 94 Tween 20 0.5% 98 Span 20 0.5% 99 Na-cholate 0.25% 96 SDS 0.25% 32 DAC 0.5% 0
IAA, lodoacetamide; NEM, N-Ethylmaleimide; SDS, Sodium dodecyl sulfate; DAC, Dimethylbenzlyalkylammonium chloride.
-
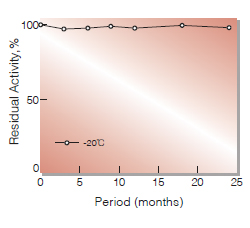
Fig.1.Stability (CRH-221)(Powder form)
(kept under dry conditions)
-
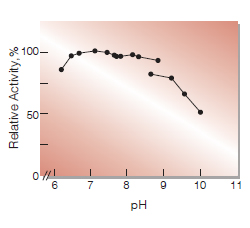
Fig.2. pH-Activity
37℃ 10min-reaction in 50mM buffer solution:pH6.2-7.8,K-phosphate; pH7.7-8.9,Tris-HCI;pH9.2-10.0, Glycine-NaOH
-
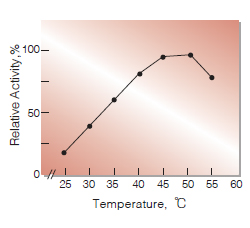
Fig.3. Temperature activity
10-min reaction in 50mM K-phosphate buffer, pH7.5
-
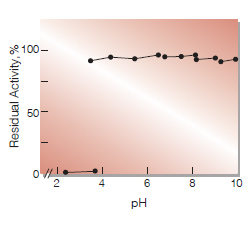
Fig.4. pH-Stability
25℃ 16hr-treatment wiht 50mM buffer solution:pH2.4-3.4,Glycine-HCI; pH3.5-6.2,Acetate; pH6.5-7.9,K-Phosphate;pH7.9-8.8,Tris-HCI;pH9.0-9.7, Glycine-NaOH
-
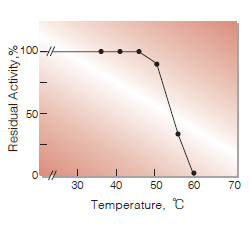
Fig.5.Thermal stability
30min-treatment with 50mM K-phosphate buffer, pH7.5
活性測定法(Japanese)
1. 原理

生成した尿素のp-ジメチルアミノベンズアルデヒド(DAB)との縮合(Ehrich反応)生成物(黄色色素)を比色定量する。
2.定義
下記条件下で1分間に1マイクロモルの黄色色素を生成する酵素量を1単位(U)とする。
3.試薬
- 0.1Mクレアチン溶液〔1.49gのクレアチン(ナカライテスク製)を50mMリン酸緩衝液 pH7.5に溶解し,100mlとする〕(用時調製)
- DAB溶液(2.0gのp-ジメチルアミノベンズアルデヒドを100mlのジメチルスルホキシドに溶解させた後,濃塩酸15mlを加える)
酵素溶液:酵素標品を予め氷冷した50mMリン酸緩衝液,pH7.5で溶解し,分析直前に同緩衝液で2.0~3.0U/mlに希釈する。
4.手順
1.試験管に基質溶液(A)1.0mlを採り,37℃で約5分間予備加温する。
2.酵素溶液0.1mlを加え,反応を開始する。
3.37℃で正確に10分間反応させた後,DAB溶液(B)2.0mlを加えて反応を停止させる。
4.25℃で20分間放置後,435nmにおける吸光度を測定する (ODtest)。
5.盲検は基質溶液(A)1.0mlを37℃で10分間放置後,DAB溶液(B)2.0mlを加えて混和し,次いで酵素溶液0.1mlを加えて調製する。以下同様に25℃で20分間放置後吸光度を測定する(ODblank)。
5.計算式
U/ml =
-
ΔOD(OD test-OD blank)×3.1(ml)×希釈倍率
0.321×1.0×10(分)×0.1(ml)
| = ΔOD×9.65×希釈倍率 | |
| U/mg | =U/ml×1/C |
| 0.321 | : 黄色色素のミリモル分子吸光係数(cm2/micromole) |
| 1.0 | : 光路長(cm) |
| C | : 溶解時の酵素濃度(c mg/ml) |
CONTACT
お問い合わせ-
各種製品に関するご質問・ご相談はこちらよりお問い合わせください。
